May 19, 2020 Particularly, the one we’re looking at is GSnap. Now, although it’s free of charge, it’s only available for Windows users. And knowing that it’s a great and useful plugin, it comes as a real bummer for Mac users. But don’t worry, as we’ve decided to look into this matter and bring you the list of the best GSnap alternatives for Mac. GSNAP is designed to align short reads from NGS data and allow detection of short and long range splicing de novo or with a database of know juctions. In addtion, SNP-tolerant alignements and alignments of reads obtained from bisulfite treated DNA are implemented. There may be multiple versions of gmap-gsnap available.
Name: GSnap
Category: Auto-tune
Developer: GVST
Date Added: September 27, 2014
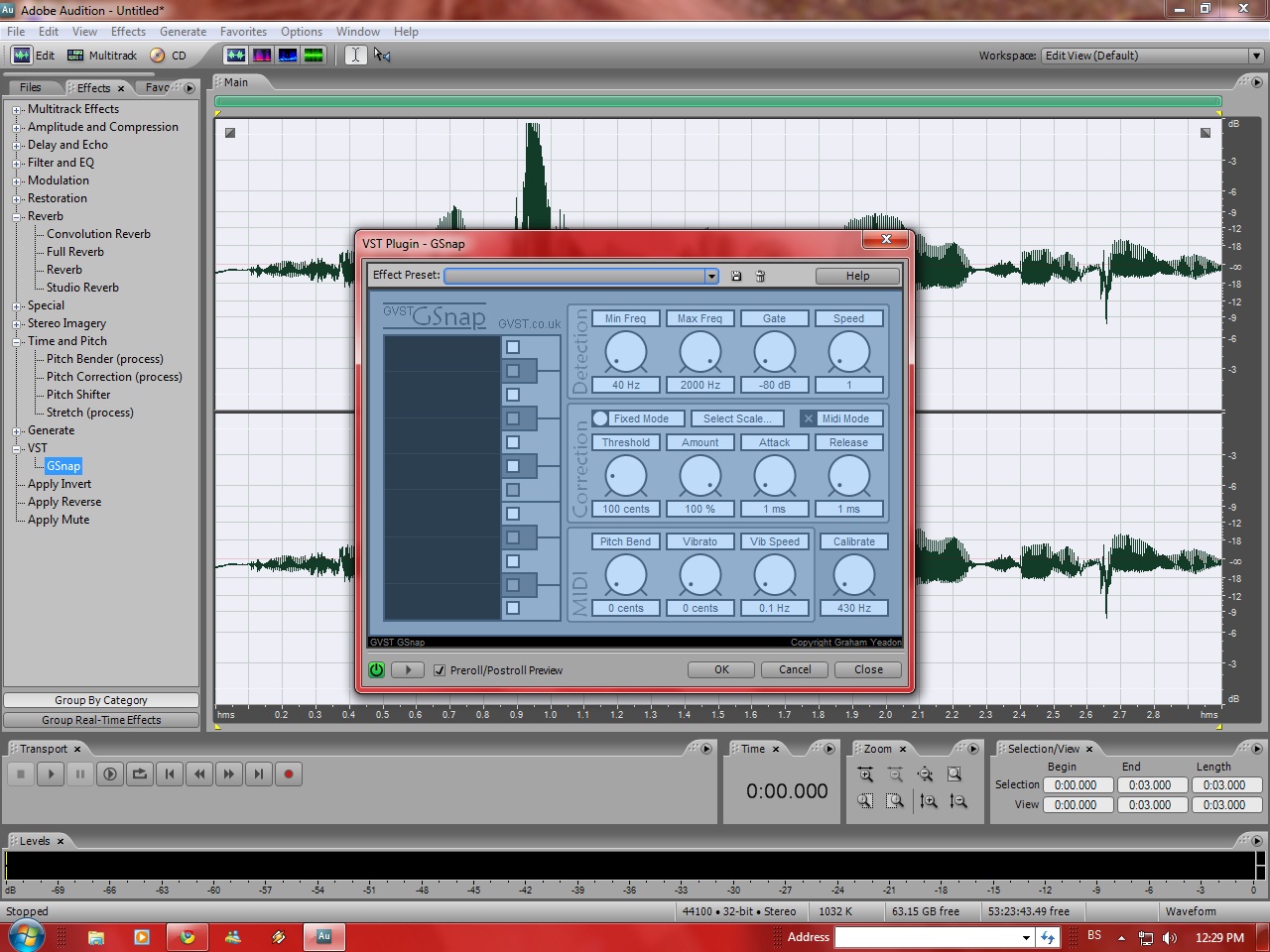
GSnap is a pitch-correction tool or auto-tune effect. This audio plugin can be used subtly to correct the pitch of a vocal, or, with more extreme settings, to create a robot-voice effect (T-Pain or Cher effect).
GSnap have two modes, Fixed mode (snap to selected notes) and MIDI mode (snap to MIDI).
Fixed mode allow us to correct the pitch based on a key and music scale, and MIDI mode used to change pitch to MIDI note, that sent either by a controller being played in real-time, or by a pre-programmed MIDI sequence. Note: For this to work, your sequencer must be set up so that GSnap can receive both audio and MIDI messages.
GSnap have two modes, Fixed mode (snap to selected notes) and MIDI mode (snap to MIDI).
Fixed mode allow us to correct the pitch based on a key and music scale, and MIDI mode used to change pitch to MIDI note, that sent either by a controller being played in real-time, or by a pre-programmed MIDI sequence. Note: For this to work, your sequencer must be set up so that GSnap can receive both audio and MIDI messages.
Support GVST
If you have found this plugin useful, please consider making a donation.GSnap is a vst instruments plugins developed by GVST , a free Auto-tune VST plugins that you can use on any VST Compatible hosts such as Steinberg Cubase, Nuendo, Wavelab, FL Studio/Fruityloops, Ableton Live, Adobe Audition, LMMS, Reaper, SONAR, Mixcraft, Acid Pro, etc.
For more information about GSnap please visit Developer Website
Description
GMAP is a tools for rapidly and accurately mapping and aligning cDNAsequences to genomic sequences.
GSNAP is designed to align short reads from NGS data and allow detection ofshort and long range splicing de novo or with a database of know juctions. Inaddtion, SNP-tolerant alignements and alignments of reads obtained frombisulfite treated DNA are implemented.
There may be multiple versions of gmap-gsnap available. An easy way of selecting theversion is to use modules. To see the modulesavailable, type
To select a module use
where
[version] is the version of choice.gmap-gsnap is a multithreaded application. Make sure to match thenumber of cpus requested with the number of threads.
Environment variables set
- $PATH
References
- Thomas D. Wu and Colin K. Watanabe. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 2005, 21:1859-1875. Pubmed | PMC] | Journal
- Thomas D. Wu and Serban Nacu. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 2010, 26:873-881. Pubmed | PMC | Journal
Documentation
Starting with release 2014-04-01 gmap/gsnap switched to a new index format.Indices are stored under
/fdb/gmap_post20140401/[organism]/[source]/[build]/Sequence/gmap-gsnap[organism]is the specific organism of interest (Gallus_gallus, Rattus_norvegicus, etc.)[source]is the source for the sequence (NCBI, Ensembl, UCSC)[build]is the specific genome build of interest (hg19, build37.2, GRCh37)
in a structure analogous to the igenomes package. In addition, commonly usedgenomes are also available at
/fdb/gmap_post20140401/common_genomes. This path is also thedefault path gmap and gsnap use to look for genome files, so a particular genomeis available there it is sufficient to provide the genome name with -dgenome_name to any of the tools without also specifying the genomedirectory with -D.Performance considerations
Gsnap throughput was measured using different numbers of threads with anENCODE data set (100nt single ended RNA-Seq, mouse Purkinje cells; accession ENCFF220BOE; biosample ENCSR758TAD). Discovery of novel splice sites was disabled but knownsplice sites were provided as input. The input file was compressed and storedon local scratch space. The output was saved unmodified on local scratch space. Eachprocess was allocated 30GB of memory. The exact command used was
Reads aligned per second per thread for different versions of gsnap with different numbers of threads.
The lower throughput of version 2015-05-01 with default settings is largely the result of a change in the default for one setting (
--end-detail) from medium to high which has the effect of improving alignments in the small number of cases where the distal end of a splice or indel contains a mismatch. Changing this version back to medium recovers most the loss in throughput. It is therefore largely a tradeoff between throughput and sensitivity. As of version 2016-06-30, the default reverted to medium.Most benchmarks were run on the phase1 norm nodes (x2650; 10g). Benchmarks ending in x2695 were run on the newer phase2 norm nodes (x2695; ibfdr).
Gsnap scales well with the number of threads all the way to 32, though the mostefficient use is probably around 24 threads with this input data set. Note that gsnap does not start any extra processes or threads, so
--nthreads=$SLURM_CPUS_PER_TASK is appropriate unless the output ispiped into another process such as samtools. In that case the number of threads usedby the second process has to be subtracted from the number of threads given togsnap.Performance was comparable when input and output were kept on the shared storagespace (
/data).Gmap and gsnap are multi-threaded applications. The thread count isdetermined with
-t / --nthreads and defaults to 1. Please do notrun with more than 2 threads on the interactive helix system.Gsnap Mac Plugin
Map and align a single cDNA to the mouse genome. Note that the mouse genomeis in common genomes, so no path has to be provided.
Map and align single end RNA-Seq data from encode to the mouse genome withdatabase of known splice junctions. Supress search for novel juctions:
A number of other small tools are available as part of the package.
Batch job on Biowulf
Gsnap Alternative Mac
Batch scripts should use the
$SLURM_CPUS_PER_TASK environmentvariable to determine the number of threads to use. This variable is set byslurm according to the number of requested cores. An example script for gsnapmight look like the following:The script is submitted to the batch system requesting, for example, 8 coresand 25GB of memory:
Set up a command file for two jobs, each aligning single ended reads. Notethat swarm allows line continuations.
Create the known splice site database as shown above and start the jobs with swarm
Interactive job on Biowulf
Interactive work requiring significant resources should be carried out oninteractive compute nodes, not on the head node or helix. Interactive nodes areallocated with
sinteractive. For example, to request a 8 coreinteractive job with 25GB memory:Gvst Gsnap
On the interactive node, gmap and gsnap are then used essentially as decribedabove.